Metagenomics is so easy to understand, right? Scientists just go out and get DNA sequences from…stuff…in the environment. And then they answer lots of questions, like….um…
Yeah sometimes I’m lost too. In metagenomics, researchers collect ocean water or soil samples and sequence random bits of DNA from whatever blob of gunk they collect–they end up with little snippets from all the genomes of the species present in that sample. But understanding metagenomics is like understanding human genomics. It’s a lot more complex and nuanced than you think, even if the overarching concept seems easy to understand. Of COURSE we want to sequence the Human Genome, just like we’re obsessed with peering into genomes of the species that live in seawater (seawater is easy to collect and we know lots of things live there; its the low-hanging fruit of marine genomics). But scientists are often asking very specific questions under these broad banners–and the questions that fall under a category such as “metagenomics” might be surprising.
The Eisen lab is currently hosting Matthew Haggerty, a visiting grad student from Elizabeth Dinsdale’s lab at San Diego State University. Last week he presented his research at lab meeting, discussing different kinds of scientific stories he’s trying to find in marine metagenomic datasets.
Matt is working with public DNA datasets collected as part of the Global Ocean Survey (= scientists who sailed around the world in a yacht and collected buckets of seawater. How can I get this job?).
The Global Ocean Survey (GOS) data has been a keystone dataset for marine science. Collected way, way back in the dark ages of genomics (2007), people have continued to look deeper and re-analyze these datasets to get the low down on ocean biology. Matt is using GOS data to look at some super cool, but super sekrit questions…at least his questions are sekrit until he gets them published. Grad students gots to gets the manuscripts peer-reviewed!
But Matt did present some pretty awesome metagenomics factoids that made me take notes in the form of iPad doodles (thank you, Notability app!).
FACT 1:
Coral reefs that are stressed or in decline show a particular increase in pathogenic bacteria and virulence genes (Ainsworth et al. 2010). Ainsworth et al. note that metagenomic studies have shown “a dramatic difference in microbial metabolic function indicated that proximity to human populations and local disturbance significantly influenced microbial diversity and metabolism and as a result influenced coral reef health.” The coral reef story is complex (involving interactions between corals, algae, bacteria, and viruses) and still being investigated, but metagenomics is now offering an unprecedented look into how coral reefs function and stay healthy (or become not so healthy).
FACT 2:
Metagenomics also lets us look at nutrient cycling in the oceans, and study how the limited availability of certain elements can control the metabolism of microbial communities. For example, Toulza et al. (2012) used that handy Global Ocean Survey data to look at genes for iron uptake in ocean microbes. Why? Because, they say, “despite the lack of knowledge of iron uptake mechanisms in most marine microorganisms, our approach provides insights into how the iron metabolic pathways of microbial communities may vary with seawater iron concentrations.” In other words, metagenomics allows us to understand how nutrients get cycled in the ocean, and how microbial cells react to different levels of nutrient availability–without ever having to know anything about any individual species.
FACT 3:
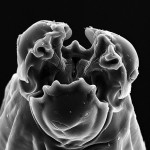
Yes, metagenomics can even involve the study of oral slime in fish–something that Matt is very interested in (although based on the number of underwater pictures, I think he may be using this question as an excuse to go diving in exotic places. Right, Matt? :P ). Parrotfish are famous for their teeth, but less famous for the green slime that grows on their teeth (see below photo). It’s pretty gross, so now you won’t forget it. I couldn’t find any peer-reviewed information about this disgusting green tooth scum, which means that the field of “parrotfish oral microbiome” needs some attention. How many species live in this slime? Does it affect parrotfish health? Do we see any cool adaptations in these cyanobacterial genomes? These questions are unanswered for now, but anyone has more info please let us know!

References:
Ainsworth TD, Thurber RV, Gates RD (2010) The future of coral reefs: a microbial perspective. Trends in Ecology & Evolution, 25(4):233–40.
Toulza E, Tagliabue A, Blain S, Piganeau G (2012) Analysis of the Global Ocean Sampling (GOS) Project for Trends in Iron Uptake by Surface Ocean Microbes. PLoS ONE, 7(2): e30931. doi:10.1371/journal.pone.0030931









Great job Matt! (and great article Holly) I’m glad you have been having fun in Davis, now come back to San Diego…. So we can hang out.
Hi,
I just spotted your article by chance, very clearly written, and thank you for the citation of our work on the microbial black box !