Today marks the long-awaited release of the Daphnia pulex genome, published today in Science. Why is this such a momentous occasion? Well first of all, there are four people from my lab whose names are on the paper, so I will probably get free beer at some point today to celebrate (score!). But more importantly, Daphnia represents the only ‘model’ organism where we even have a vague idea of its ecology and life-history. I dare you to try and find C. elegans in the soil somewhere–its just not gonna happen . Yet this worm was the first multicellular organism to have its entire genome sequenced, and there is still a veritable battilion of scientists feverishly working to understand its genome. (All C. elegans work these days is done on strains that have been in culture for years, and the orginal worms were found randomly living in piles of compost and garbage.)
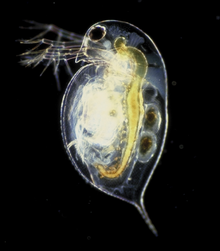
But for Daphnia, a.k.a. the ‘water flea’, we have a REALLY good idea of what this species actually DOES in the wild. These animals are small (0.2-0.5 mm), free-swimming crustaceans that are fairly ubiquitous in freshwater. They can tolerate a huge range of aquatic conditions, and their sensitivity to toxins makes them an ideal indicator species for assessing the health of an environment. They also don’t live long (big plus for growing them in the lab), with their life cycle (reproducing by cloning vs. sexually) and anatomical features (cool anti-predator defenses) affected by the environment in which they live:

Armed with this information, a bunch of scientists said “Hey, lets grow this weird animal in a whole bunch of different toxins and water conditions, sequence the genome, and then see how gene expression changes across the different treatments”. Which is exactly what the Science paper is about. It turns out that Daphnia pulex has a tiny genome containing only 200 million base pairs (compared to 3 billion in humans), yet has a whopping 31,907 genes within it (that’s more than humans, who have 20-25,000 genes!). More than a third of Daphnia genes are unique to this species. The high number of genes is a result of duplication, where the genome contains multiple copies of similar genes–this explains how Daphnia can adapt to a wide range of environmental conditions and stress. The rationale for maintaining duplicate genes is kind of like my black shoe collection. Black shoes go with everything, but not all pairs are interchangeable: sky-high black stilettos are appropriate for a martini bar (but not for work!) while my black granny pumps are reserved for job interviews (and I wouldn’t be caught dead wearing them on a date).
So as genes have duplicated in Daphnia, the two new copies a given gene (called paralogs) have acquired slightly different characteristics. The animal’s environment may then determine which gene gets expressed–i.e. which gene allows it to adapt better to its surrounding ecosystem. Gene duplication isn’t this cut and dry–sometimes duplicated genes are lost and become non-functional, or the expression of duplicated genes diverges wildly as mutations accumulate over time–but its pretty cool nonetheless. Now that the Daphnia genome is published, we have a better understanding of how an animal’s environment impacts cellular machinery.
Reference:
John K. Colbourne, et al. (2011) The Ecoresponsive Genome of Daphnia pulex. Science 331: 555-561. DOI: 10.1126/science.1197761





Good points – key genetic work on a species known in its ecological context – takes me back to looking at zooplankton grazing dynamics in the tropics. I’ve still got Korinek’s guide to African inland Cladocera somewhere…
I love your analogy of duplicated genes to shoes! That analogy would never have occurred to me, but I can see how effective it would be for my wife and daughter (and the female students in my freshman bio class).
My second grade class was “wowed” by the news that this “rice” sized crustacean can actually transform itself into something resembling a sword and shield when confronted by a predator! It’s like one of their beloved pokemans coming alive! Thank you as I work to make science fascinating to my students…
I was just talking to a worm researcher yesterday about how little is know about their ecology. Such a ridiculous disparity between what they know about its genetics and development, versus the animal’s actual lifestyle.
I’m also working on a post about the Daphnia genome, but related to the visual system. Hopefully I can get that done later today.