
Editor’s Note: Genomic Repairman is a friend I’ve gotten to know through Twitter (@genrepair). He is a semi-cultured, good-natured graduate student in biomedical sciences who escaped out of the deep south and now focuses on using genetics and biochemistry to elucidate DNA repair in cells. He blogs at Tales From a Genomic Repairman. I asked him to guest post here which he graciously accepted and offered this excellent post on how studying fish has helped us to understand cancer. Enjoy!
————————————————————————–
Let me begin with an admission of ignorance, I am not a marine biologist, I do not have a great fundamental understanding of fish or the life aquatic. But I do not let little things like an utter complete lack of knowledge or not having really worked in marine science hold me back from making a post. So my aim is to take a stab at this topic of fish by talking about DNA which I know a great deal about and study how it becomes broken and then how it gets repaired.
Before we get to fish, lets amble on down to another topic, cancer, and more specifically melanoma. Cutaneous malignant melanoma (CMM) is an aggressive form of skin cancer whose incidence seems to be increasing worldwide (1). It is understood that sun exposure is the predominant environmental cause of this cancer but there is likely to be some hereditary disposition for CMM also (2-4).
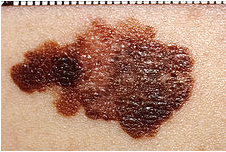
Cancer research tends to focus on mammalian models for obvious reasons (comparable cell lineage, physiologic similarity to humans, and utility of generating mutants in the case of mice). Non-mammalian models, such as fish, are of great value as well as they are some of the oldest models of cancer and have been proven to be effective (5). Relatively short breeding cycles, large progeny, and low cost of use provide a substantial advantage to their use as models of disease.
Tumor formation in the fish genus Xiphophorus has been well established. Way way back, since after the first great war (that’s WWI to all you that got D’s in history), certain hybrids of the genus (platyfish and swordtails) developed malignant tumors of pigment cells that were classified as melanomas (6). Interestingly enough melanomas from Xiphophorus can be xenografted (a process of transfering tissue between different hosts) onto nude mice and grown. More interestingly, it still maintains fish antigen expression (7). The melanoma-specific cancer gene (e.g. oncogene) locus, known as Tu (as in tumor), is repressed by the R (regulator) locus on another chromosome in fish and selective breeding can be undertaken (8, 9). Through a classical crossing experiment you can develop a variety of pigmented phenotypes by selecting for Tu and breeding R out of the hybrids. This is really dry and if you want more detail I would be more than happy to talk to you about it, but lets face it breeding strategies are quite boring and better left to personal discussions than blog posts.
So what is this Tu? Cloning and genetic disruption experiments revealed that the gene at the Tu locus is EGFR-related and is responsable for melanoma formation (10, 11). The gene Xmrk (Xiphophorus melanoma receptor kinase) arose from a tandem duplication event of Xiphophorus EGFR and was found to be highly expressed (i.e. “turned on”) in transformed pigment cells of certain hybrids (12). So what is Xmrk doing in the cell? Xmrk is working through the classic Ras-Raf-MAPK pathway to stimulate cell proliferation as well as activating STAT5, which promotes anti-apoptotic (apoptosis=cell death) signaling as well as in addition to proliferation.
Enough about Xmrk, what is R? Well we only kind of have an idea and its not too solid. While genetic evidence points to the CDKN2AB gene, a homologue of cyclin-dependant kinase inhibitor p16, to be the candidate tumor suppressor, the functional evidence so far is unclear (8). Therefore this remains an area of intense focus for fish researchers. Similarly further characterization of tumor modifier genes still needs to be sorted out as well. Recently, a new fish melanoma model has been reported in the literature. The Schartl lab used the medaka fish (Oryzias latipes) to create a transgenic Xmrk fish that results in a highly invasive and more metastatic melanoma, which may be more suitable model of human melanoma (13).
Before leaving you guys, lets talk about another possible benefit of using fish as a model of melanoma. Small molecule inhibitor screens are all the rage these days and these fish could be integral in finding chemicals that interfere with different aspects of melanocyte biology: pigmentation, migration, and cell survival. This in vivo testing model involves placing embryos in well plates and treating with specific chemicals in the water, creating a cheap and rapid in vivo testing system for small molecules.
Marine science research is important and can have implications on human health as well. This is but one example how someone studying fish noticed a phenomena that had suitable implications for a human disease. So go study fish, who knows, your research might be helpful in fighting cancer one day. So folks this is where I’m going to stop the car and let you out. I hope this has been informative and not too dry. Once again, my focus is fish, so I may have left some gaps but if you have questions I’ll try to fill them in as best as possible.
References:
1. Linos E, Swetter SM, Cockburn MG, Colditz GA, Clarke CA. Increasing burden of melanoma in the United States. J Invest Dermatol. 2009;129:1666-74.
2. Bishop JN, Harland M, Randerson-Moor J, Bishop DT. Management of familial melanoma. Lancet Oncol. 2007;8:46-54.
3. Chin L, Garraway LA, Fisher DE. Malignant melanoma: genetics and therapeutics in the genomic era. Genes Dev. 2006;20:2149-82.
4. Rivers JK. Is there more than one road to melanoma? Lancet. 2004;363:728-30.
5. Friend SH. Genetic models for studying cancer susceptibility. Science. 1993;259:774-5.
6. Gordon M. The Genetics of a Viviparous Top-Minnow Platypoecilus; the Inheritance of Two Kinds of Melanophores. Genetics. 1927;12:253-83.
7. Schartl M, Peter RU. Progressive growth of fish tumors after transplantation into thymus-aplastic (nu/nu) mice. Cancer Res. 1988;48:741-4.
8. Kazianis S, Gutbrod H, Nairn RS, McEntire BB, Della Coletta L, Walter RB, et al. Localization of a CDKN2 gene in linkage group V of Xiphophorus fishes defines it as a candidate for the DIFF tumor suppressor. Genes Chromosomes Cancer. 1998;22:210-20.
9. Leroi AM, Koufopanou V, Burt A. Cancer selection. Nat Rev Cancer. 2003;3:226-31.
10. Schartl M, Hornung U, Gutbrod H, Volff JN, Wittbrodt J. Melanoma loss-of-function mutants in Xiphophorus caused by Xmrk-oncogene deletion and gene disruption by a transposable element. Genetics. 1999;153:1385-94.
11. Wittbrodt J, Adam D, Malitschek B, Maueler W, Raulf F, Telling A, et al. Novel putative receptor tyrosine kinase encoded by the melanoma-inducing Tu locus in Xiphophorus. Nature. 1989;341:415-21.
12. Gomez A, Volff JN, Hornung U, Schartl M, Wellbrock C. Identification of a second egfr gene in Xiphophorus uncovers an expansion of the epidermal growth factor receptor family in fish. Mol Biol Evol. 2004;21:266-75.
13. Schartl M, Wilde B, Laisney JA, Taniguchi Y, Takeda S, Meierjohann S. A mutated EGFR is sufficient to induce malignant melanoma with genetic background-dependent histopathologies. J Invest Dermatol. 2010;130:249-58.





Great job, GR! Fish are great models for all sorts of vertebrate biology, and you did a great job highlighting the cancer aspect.
I also like this:
“but lets face it breeding strategies are quite boring and better left to personal discussions than blog posts.”
hahahaha!! It would be quite a different post if you were focused on breeding strategies, eh?! ;-)